I have a following survival data
library(survival)
data(pbc)
#model to be plotted and analyzed, convert time to years
fit <- survfit(Surv(time/365.25, status) ~ edema, data = pbc)
#visualize overall survival Kaplan-Meier curve
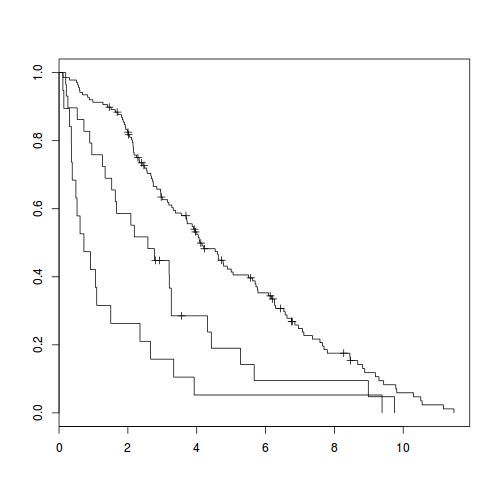
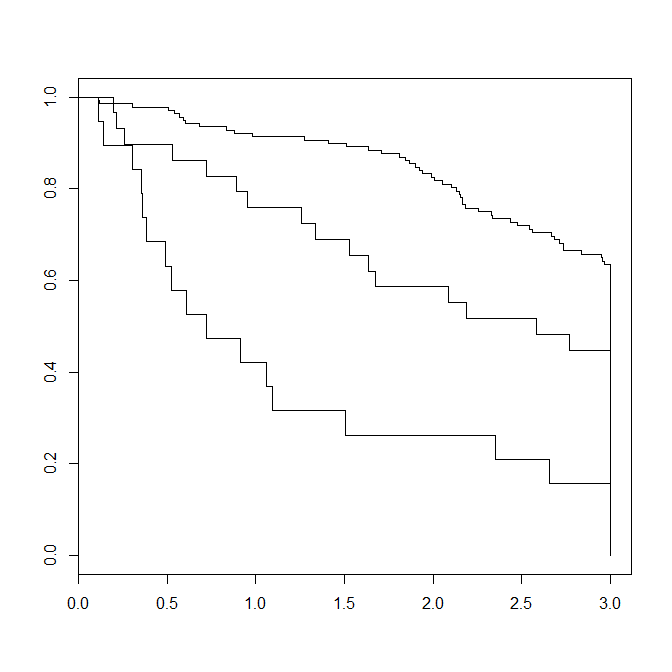
plot(fit)
Here is how the resulting Kaplan-Meier plot looks like
I am further calculating survival at 1, 2, 3 years in this manner:
> summary(fit,times=c(1,2,3))
Call: survfit(formula = Surv(time/365.25, status) ~ edema, data = pbc)
232 observations deleted due to missingness
edema=0
time n.risk n.event survival std.err lower 95% CI upper 95% CI
1 126 12 0.913 0.0240 0.867 0.961
2 112 12 0.825 0.0325 0.764 0.891
3 80 26 0.627 0.0420 0.550 0.714
edema=0.5
time n.risk n.event survival std.err lower 95% CI upper 95% CI
1 22 7 0.759 0.0795 0.618 0.932
2 17 5 0.586 0.0915 0.432 0.796
3 11 4 0.448 0.0923 0.299 0.671
edema=1
time n.risk n.event survival std.err lower 95% CI upper 95% CI
1 8 11 0.421 0.1133 0.2485 0.713
2 5 3 0.263 0.1010 0.1240 0.558
3 3 2 0.158 0.0837 0.0559 0.446
As you can see, the resulting output shows me 95% confidence intervals between different levels of edema but no actual p values. Whether confidence intervals overlap or not, I still get a pretty good idea whether survival at these time points are signifiantly different or not, but I would like to have exact p values. How can I do that?