In order to plot half densities, I am using the function described in this post: Split violin plot with ggplot2
However, when I want to draw the quantiles on the densities, like on a normal geom_violin() or geom_boxplot(), I obtain an error message.
I would also be interested in adding the number of observations above each half density.
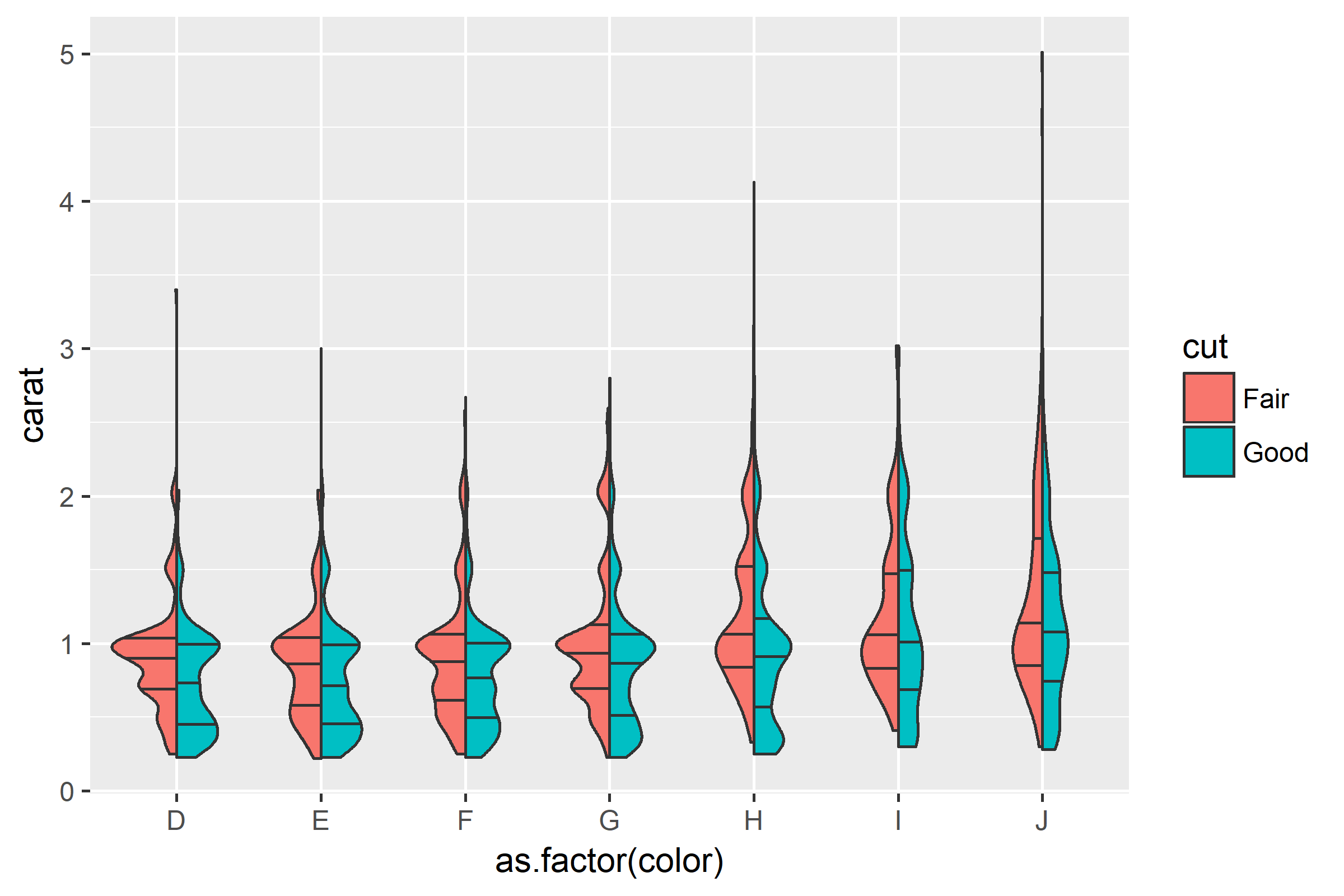
Here is an example of what I would like to obtain:
data("diamonds")
library(ggplot2)
# Function described in a previous post
GeomSplitViolin <- ggproto("GeomSplitViolin", GeomViolin, draw_group = function(self, data, ..., draw_quantiles = NULL){
data <- transform(data, xminv = x - violinwidth * (x - xmin), xmaxv = x + violinwidth * (xmax - x))
grp <- data[1,'group']
newdata <- plyr::arrange(transform(data, x = if(grp%%2==1) xminv else xmaxv), if(grp%%2==1) y else -y)
newdata <- rbind(newdata[1, ], newdata, newdata[nrow(newdata), ], newdata[1, ])
newdata[c(1,nrow(newdata)-1,nrow(newdata)), 'x'] <- round(newdata[1, 'x'])
if (length(draw_quantiles) > 0 & !scales::zero_range(range(data$y))) {
stopifnot(all(draw_quantiles >= 0), all(draw_quantiles <=
1))
quantiles <- create_quantile_segment_frame(data, draw_quantiles)
aesthetics <- data[rep(1, nrow(quantiles)), setdiff(names(data), c("x", "y")), drop = FALSE]
aesthetics$alpha <- rep(1, nrow(quantiles))
both <- cbind(quantiles, aesthetics)
quantile_grob <- GeomPath$draw_panel(both, ...)
ggplot2:::ggname("geom_split_violin", grobTree(GeomPolygon$draw_panel(newdata, ...), quantile_grob))
}
else {
ggplot2:::ggname("geom_split_violin", GeomPolygon$draw_panel(newdata, ...))
}
})
geom_split_violin <- function (mapping = NULL, data = NULL, stat = "ydensity", position = "identity", ..., draw_quantiles = NULL, trim = TRUE, scale = "area", na.rm = FALSE, show.legend = NA, inherit.aes = TRUE) {
layer(data = data, mapping = mapping, stat = stat, geom = GeomSplitViolin, position = position, show.legend = show.legend, inherit.aes = inherit.aes, params = list(trim = trim, scale = scale, draw_quantiles = draw_quantiles, na.rm = na.rm, ...))
}
tmp <- diamonds[which(diamonds$cut %in% c("Fair", "Good")), ]
# Obtained plot
ggplot(tmp, aes(as.factor(color), carat, fill = cut)) +
geom_split_violin()
# Error due to internal functions (interleave, ...)
ggplot(tmp, aes(as.factor(color), carat, fill = cut)) +
geom_split_violin(draw_quantiles = 0.5)
# Function to return number of observation
give_n = function(x, y_up = y_upper) {
data.frame(y = y_up * 1.06,
label = paste("n =", length(x))
)
}
# Code to add number of observations above each half density
new_plot = given_plot +
# Give back only length of data
stat_summary(fun.data = give_n, aes(x = as.factor(variable)), geom = "text")

ggplot2:::create_quantile_segment_frameandgrid::grobTreeinGeomSplitViolin, although the result might not be satisfactory. – Clemenciaclemency