I could really use a tip to help me plotting a decision boundary to separate to classes of data. I created some sample data (from a Gaussian distribution) via Python NumPy. In this case, every data point is a 2D coordinate, i.e., a 1 column vector consisting of 2 rows. E.g.,
[ 1
2 ]
Let's assume I have 2 classes, class1 and class2, and I created 100 data points for class1 and 100 data points for class2 via the code below (assigned to the variables x1_samples and x2_samples).
mu_vec1 = np.array([0,0])
cov_mat1 = np.array([[2,0],[0,2]])
x1_samples = np.random.multivariate_normal(mu_vec1, cov_mat1, 100)
mu_vec1 = mu_vec1.reshape(1,2).T # to 1-col vector
mu_vec2 = np.array([1,2])
cov_mat2 = np.array([[1,0],[0,1]])
x2_samples = np.random.multivariate_normal(mu_vec2, cov_mat2, 100)
mu_vec2 = mu_vec2.reshape(1,2).T
When I plot the data points for each class, it would look like this:

Now, I came up with an equation for an decision boundary to separate both classes and would like to add it to the plot. However, I am not really sure how I can plot this function:
def decision_boundary(x_vec, mu_vec1, mu_vec2):
g1 = (x_vec-mu_vec1).T.dot((x_vec-mu_vec1))
g2 = 2*( (x_vec-mu_vec2).T.dot((x_vec-mu_vec2)) )
return g1 - g2
I would really appreciate any help!
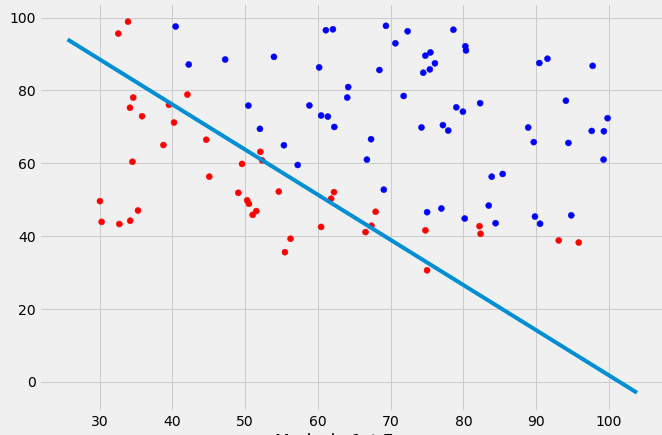
EDIT: Intuitively (If I did my math right) I would expect the decision boundary to look somewhat like this red line when I plot the function...















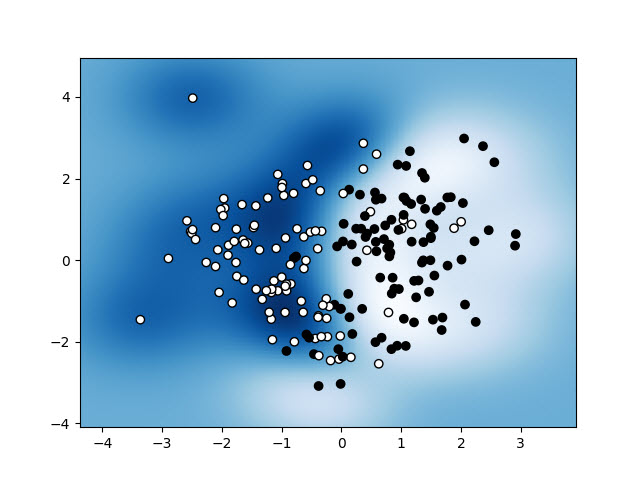
x_vecin thedecision_boundaryfunction supposed to be? Are you just trying to plot a line separating the two classes? – Radbundecision_boundary. It's easiest to just evaluate the function on a regular grid and contour the result. Hopefully that gets you pointed in the right direction! – Dragelin