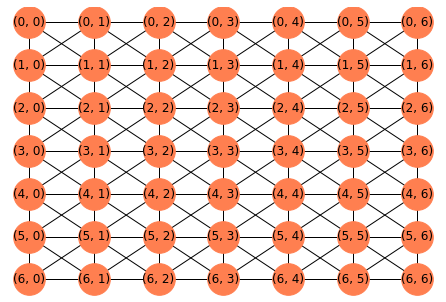
I have a gray scale image with values between 0 (black) and white (255). I have a target matrix of the same size as the gray scale image. I need to start at a random pixel in the gray scale image and traverse through the image one pixel at a time (in a depth-first search manner), copying its value to the corresponding location in the target matrix. I obviously need to do this only for the non-white pixels. How can I do this? I thought that I could get the connected components of the gray scale image and traverse each pixel one by one, but I couldn't find any suitable implementation of connected components. Any ideas?
For example, if my gray scale image is:
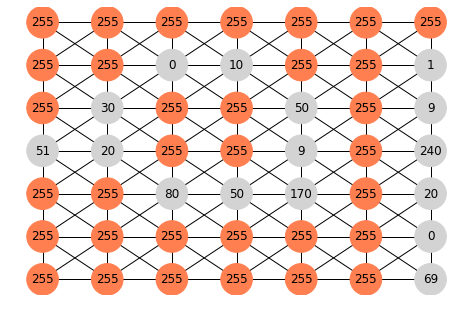
[[255,255,255,255,255,255,255]
[255,255, 0 ,10 ,255,255, 1 ]
[255,30 ,255,255,50 ,255, 9 ]
[51 ,20 ,255,255, 9 ,255,240]
[255,255,80 ,50 ,170,255, 20]
[255,255,255,255,255,255, 0 ]
[255,255,255,255,255,255, 69]]
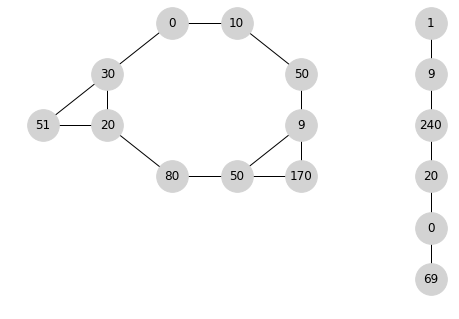
Then a possible traversal is [0,10,50,9,170,50,80,20,51,30] followed by [1,9,240,20,0,69] to give [0,10,50,9,170,50,80,20,51,30,1,9,240,20,0,69]. The order between the different objects doesn't matter.
Other possible traversals are:
[1,9,240,20,0,69,0,10,50,9,170,50,80,20,51,30] or [1,9,240,20,0,69,0,10,50,9,170,50,80,20,30,51] or
[1,9,240,20,0,69,10,50,9,170,50,80,20,30,0,51]
etc.

dfs? – Ralli01? Have you managed to at least write the code that splits the0from the1? Where's that please? – Rolfston240,9,1,20,0,69,0,10,50,9,170,50,80,20,51,30– Spicynetworkxhas a pretty good one but performance is really slow and unsatisfactory for me. Then I have found a solution usingigraph. I'm going to describe it asap. – Daile