I would assume here that your problem can be simplified by assuming that your function has to be close to another function (e.g. the center of the tube) with the very same support points and then a certain number of discontinuities are allowed.
Then, I would implement a different discretization of function compared to the typical one that is used for L^2 norm (See for example some reference here).
Basically, in the continuous case, the L^2 norm relaxes the constrain of the two function being close everywhere, and allow it to be different on a finite number of points, called singularities
This works because there are an infinite number of points where to calculate the integral, and a finite number of points will not make a difference there.
However, since there are no continuous functions here, but only their discretization, the naive approach will not work, because any singularity will contribute potentially significantly to the final integral value.
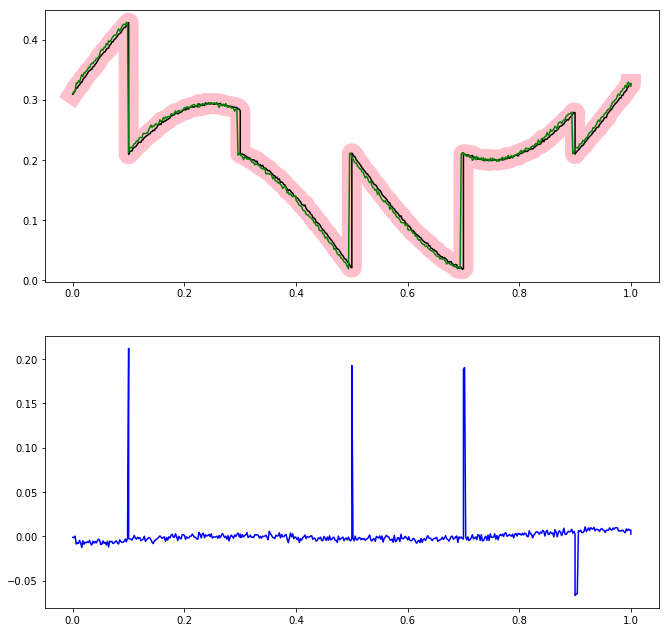
Therefore, what you could do is to perform a point by point check whether the two functions are close (within some tolerance) and allow at most num_exceptions points to be off.
import numpy as np
def is_close_except(arr1, arr2, num_exceptions=0.01, **kwargs):
# if float, calculate as percentage of number of points
if isinstance(num_exceptions, float):
num_exceptions = int(len(arr1) * num_exceptions)
num = len(arr1) - np.sum(np.isclose(arr1, arr2, **kwargs))
return num <= num_exceptions
By contrast the standard L^2 norm discretization would lead to something like this integrated (and normalized) metric:
import numpy as np
def is_close_l2(arr1, arr2, **kwargs):
norm1 = np.sum(arr1 ** 2)
norm2 = np.sum(arr2 ** 2)
norm = np.sum((arr1 - arr2) ** 2)
return np.isclose(2 * norm / (norm1 + norm2), 0.0, **kwargs)
This however will fail for arbitrarily large peaks, unless you set such a large tolerance than basically anything results as "being close".
Note that the kwargs is used if you want to specify a additional tolerance constraints to np.isclose() or other of its options.
As a test, you could run:
import numpy as np
import numpy.random
np.random.seed(0)
num = 1000
snr = 100
n_peaks = 5
x = np.linspace(-10, 10, num)
# generate ground truth
y = np.sin(x)
# distributed noise
y2 = y + np.random.random(num) / snr
# distributed noise + peaks
y3 = y + np.random.random(num) / snr
peak_positions = [np.random.randint(num) for _ in range(n_peaks)]
for i in peak_positions:
y3[i] += np.random.random() * snr
# for distributed noise, both work with a 1/snr tolerance
is_close_l2(y, y2, atol=1/snr)
# output: True
is_close_except(y, y2, atol=1/snr)
# output: True
# for peak noise, since n_peaks < num_exceptions, this works
is_close_except(y, y3, atol=1/snr)
# output: True
# and if you allow 0 exceptions, than it fails, as expected
is_close_except(y, y3, num_exceptions=0, atol=1/snr)
# output: False
# for peak noise, this fails because the contribution from the peaks
# in the integral is much larger than the contribution from the rest
is_close_l2(y, y3, atol=1/snr)
# output: False
There are other approaches to this problem involving higher mathematics (e.g. Fourier or Wavelet transforms), but I would stick to the simplest.
EDIT (updated):
However, if the working assumption does not hold or you do not like, for example because the two functions have different sampling or they are described by non-injective relations.
In that case, you can follow the center of the tube using (x, y) data and the calculate the Euclidean distance from the target (the tube center), and check that this distance is point-wise smaller than the maximum allowed (the tube size):
import numpy as np
# assume it is something with shape (N, 2) meaning (x, y)
target = ...
# assume it is something with shape (M, 2) meaning again (x, y)
trajectory = ...
# calculate the distance minimum distance between each point
# of the trajectory and the target
def is_close_trajectory(trajectory, target, max_dist):
dist = np.zeros(trajectory.shape[0])
for i in range(len(dist)):
dist[i] = np.min(np.sqrt(
(target[:, 0] - trajectory[i, 0]) ** 2 +
(target[:, 1] - trajectory[i, 1]) ** 2))
return np.all(dist < max_dist)
# same as above but faster and more memory-hungry
def is_close_trajectory2(trajectory, target, max_dist):
dist = np.min(np.sqrt(
(target[:, np.newaxis, 0] - trajectory[np.newaxis, :, 0]) ** 2 +
(target[:, np.newaxis, 1] - trajectory[np.newaxis, :, 1]) ** 2),
axis=1)
return np.all(dist < max_dist)
The price of this flexibility is that this will be a significantly slower or memory-hungry function.

L^2(orl^2) distance between the two distribution, i.e. something that will be close to zero if the two distribution differ only in at most N number of points (with N finite in the case of continuous distributions). Am I correct? – Quackery