AFAIK, there is no official implementation of partial correlation in scipy / numpy. As pointed out by @J. C. Rocamonde, the function from that stats website can be used to calculate partial correlation.
I believe here's the original source:
https://gist.github.com/fabianp/9396204419c7b638d38f
Note:
As discussed in the github page, you may want to add a column of ones to add a bias term to your fits if your data is not standardized (Judging from your data it's not).
If I'm not mistaken, it calculates partial correlation by controlling all other remaining variables in the matrix. If you just want to control one variable, you may change idx to the index of that particular variable.
Edit 1 (How to add ones + What to do with df):
If you look into the link, they have already discussed how to add ones.
To illustrate how it works, I added another way of hstack, using the given data in the link:
data_int = np.hstack((np.ones((data.shape[0],1)), data))
test1 = partial_corr(data_int)[1:, 1:]
print(test1)
# You can also add it on the right, as long as you select the correct coefficients
data_int_2 = np.hstack((data, np.ones((data.shape[0],1))))
test2 = partial_corr(data_int_2)[:-1, :-1]
print(test2)
data_std = data.copy()
data_std -= data.mean(axis=0)[np.newaxis, :]
data_std /= data.std(axis=0)[np.newaxis, :]
test3 = partial_corr(data_std)
print(test3)
Output:
[[ 1. -0.54341003 -0.14076948]
[-0.54341003 1. -0.76207595]
[-0.14076948 -0.76207595 1. ]]
[[ 1. -0.54341003 -0.14076948]
[-0.54341003 1. -0.76207595]
[-0.14076948 -0.76207595 1. ]]
[[ 1. -0.54341003 -0.14076948]
[-0.54341003 1. -0.76207595]
[-0.14076948 -0.76207595 1. ]]
And if you want to maintain the columns, easiest way is to extract the columns and put them back in after calculation:
# Assume that we have a DataFrame with columns x, y, z
data_as_df = pd.DataFrame(data, columns=['x','y','z'])
data_as_array = data_as_df.values
partial_corr_array = partial_corr(np.hstack((np.ones((data_as_array.shape[0],1)), data_as_array))
)[1:,1:]
corr_df = pd.DataFrame(partial_corr_array, columns = data_as_df.columns)
print(corr_df)
Output:
x y z
0 1.000 -0.543 -0.141
1 -0.543 1.000 -0.762
2 -0.141 -0.762 1.000
Hope it's helpful! Let me know if anything is unclear!
Edit 2:
I think the problem lies in not having constant term in each of the fits... I rewrote the code in sklearn to make it easier to add intercept:
def calculate_partial_correlation(input_df):
"""
Returns the sample linear partial correlation coefficients between pairs of variables,
controlling for all other remaining variables
Parameters
----------
input_df : array-like, shape (n, p)
Array with the different variables. Each column is taken as a variable.
Returns
-------
P : array-like, shape (p, p)
P[i, j] contains the partial correlation of input_df[:, i] and input_df[:, j]
controlling for all other remaining variables.
"""
partial_corr_matrix = np.zeros((input_df.shape[1], input_df.shape[1]));
for i, column1 in enumerate(input_df):
for j, column2 in enumerate(input_df):
control_variables = np.delete(np.arange(input_df.shape[1]), [i, j]);
if i==j:
partial_corr_matrix[i, j] = 1;
continue
data_control_variable = input_df.iloc[:, control_variables]
data_column1 = input_df[column1].values
data_column2 = input_df[column2].values
fit1 = linear_model.LinearRegression(fit_intercept=True)
fit2 = linear_model.LinearRegression(fit_intercept=True)
fit1.fit(data_control_variable, data_column1)
fit2.fit(data_control_variable, data_column2)
residual1 = data_column1 - (np.dot(data_control_variable, fit1.coef_) + fit1.intercept_)
residual2 = data_column2 - (np.dot(data_control_variable, fit2.coef_) + fit2.intercept_)
partial_corr_matrix[i,j] = stats.pearsonr(residual1, residual2)[0]
return pd.DataFrame(partial_corr_matrix, columns = input_df.columns, index = input_df.columns)
# Generating data in our minion world
test_sample = 10000;
Math_score = np.random.randint(100,600, size=test_sample) + 20 * np.random.random(size=test_sample)
Eng_score = np.random.randint(100,600, size=test_sample) - 10 * Math_score + 20 * np.random.random(size=test_sample)
Phys_score = Math_score * 5 - Eng_score + np.random.randint(100,600, size=test_sample) + 20 * np.random.random(size=test_sample)
Econ_score = np.random.randint(100,200, size=test_sample) + 20 * np.random.random(size=test_sample)
Hist_score = Econ_score + 100 * np.random.random(size=test_sample)
minions_df = pd.DataFrame(np.vstack((Math_score, Eng_score, Phys_score, Econ_score, Hist_score)).T,
columns=['Math', 'Eng', 'Phys', 'Econ', 'Hist'])
calculate_partial_correlation(minions_df)
Output:
---- ---------- ----------- ------------ ----------- ------------
Math 1 -0.322462 0.436887 0.0104036 -0.0140536
Eng -0.322462 1 -0.708277 0.00802087 -0.010939
Phys 0.436887 -0.708277 1 0.000340397 -0.000250916
Econ 0.0104036 0.00802087 0.000340397 1 0.721472
Hist -0.0140536 -0.010939 -0.000250916 0.721472 1
---- ---------- ----------- ------------ ----------- ------------
Please let me know if that's not working!

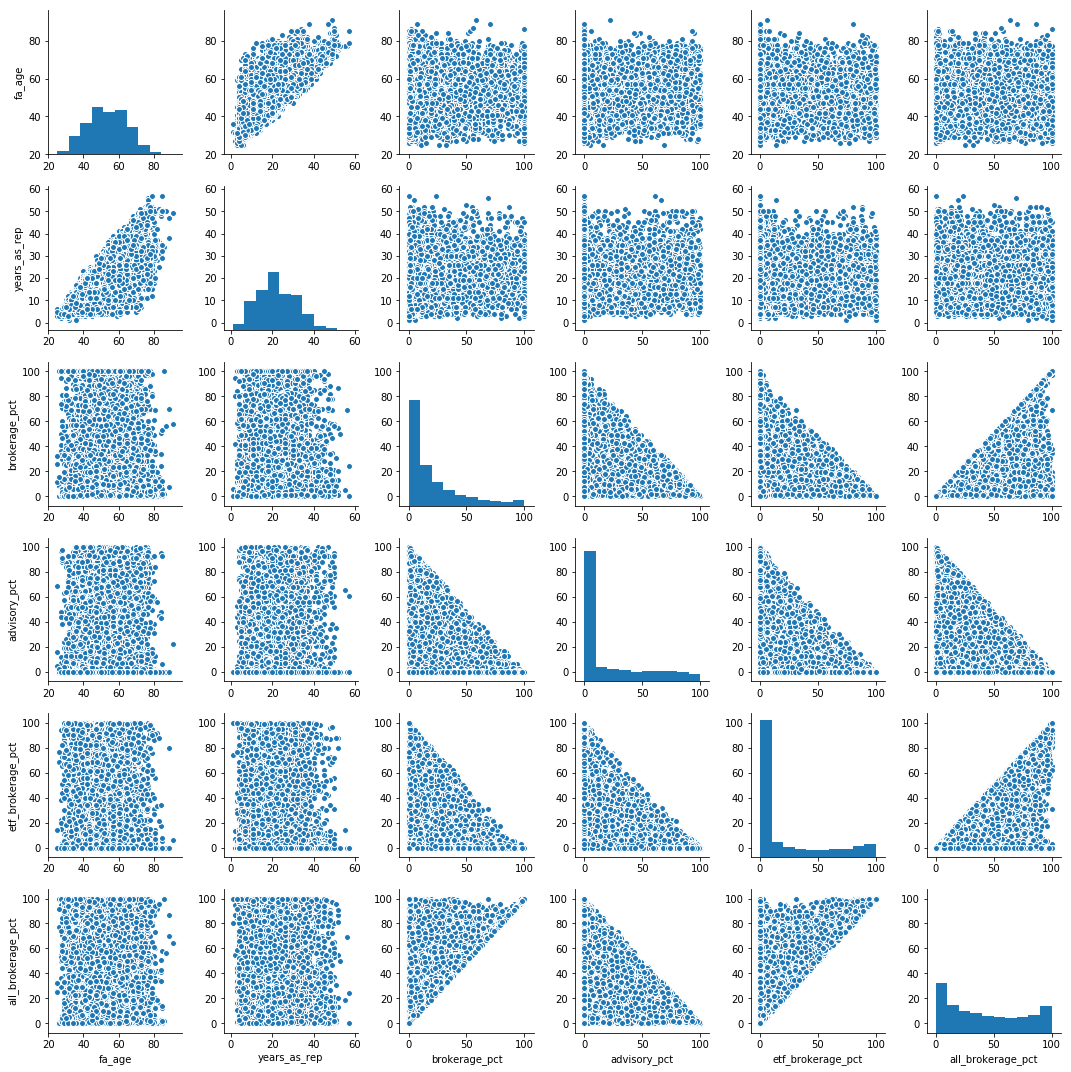
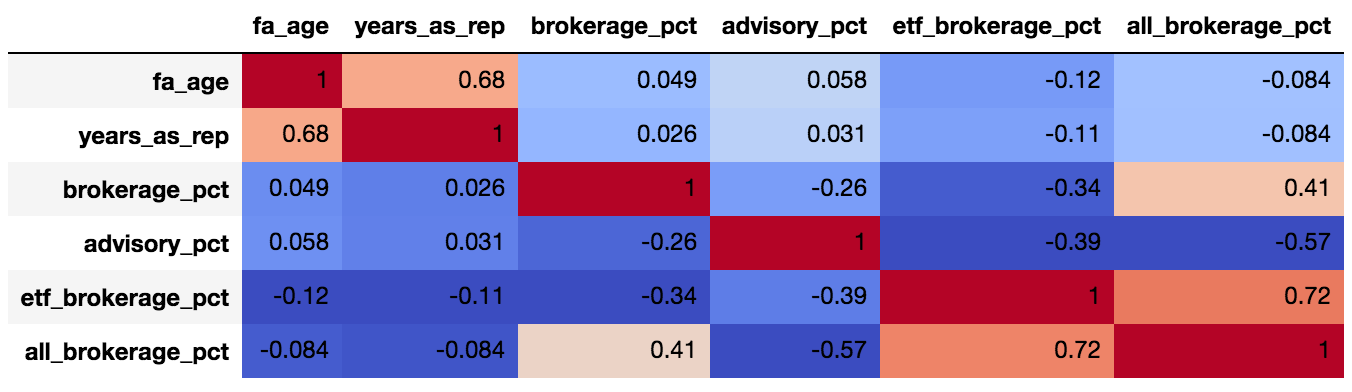
np.random.randint()your chose for the various variables? – Hoyedict = {'x1': [1, 2, 3, 4, 5], 'x2': [2, 2, 3, 4, 2], 'x3': [10, 9, 5, 4, 9], 'y' : [5.077, 32.330, 65.140, 47.270, 80.570]} data = pd.DataFrame(dict, columns=['x1', 'x2', 'x3', 'y'])– Hoye